A significant breakthrough in understanding the complexities of cancer treatment efficacy has emerged from a new study published in Nature Communications. Led by Dr. Louise Fets at the MRC Laboratory of Medical Sciences (LMS), the research team delved into the perplexing phenomenon of why the same cancer therapy can yield dramatically different outcomes across patients. Focusing on PARP inhibitors, a crucial class of targeted cancer drugs, the scientists employed advanced imaging techniques to meticulously track drug movement within ovarian tumor samples, revealing that cellular ‘recycling centers’ known as lysosomes play a pivotal and previously underappreciated role in drug distribution and overall treatment success.
The variability in patient response to cancer therapies represents one of the most formidable challenges in modern oncology. While targeted drugs have revolutionized treatment landscapes for many malignancies, a substantial proportion of patients either do not respond to initial therapy or develop resistance over time. This new research offers a compelling explanation, demonstrating that PARP inhibitors can accumulate within lysosomes, where they become temporarily trapped before being released. This dynamic interaction within individual cells fundamentally influences drug availability and, consequently, the effectiveness of the treatment.
The Evolving Landscape of Cancer Treatment and the PARP Inhibitor Revolution
For decades, cancer treatment primarily relied on surgery, radiation, and conventional chemotherapy, which indiscriminately targeted rapidly dividing cells. While often effective, these approaches frequently caused severe side effects due to damage to healthy tissues. The dawn of the 21st century heralded a new era of targeted therapies, drugs designed to specifically interfere with molecular pathways critical for cancer cell growth and survival, while minimizing harm to normal cells.
Among these advancements, PARP inhibitors have been particularly transformative, especially for patients with ovarian, breast, and prostate cancers characterized by deficiencies in DNA repair mechanisms, such as mutations in the BRCA1 or BRCA2 genes. Poly(ADP-ribose) polymerase (PARP) is a family of enzymes involved in DNA repair, specifically single-strand break repair. When PARP is inhibited, cancer cells with existing DNA repair deficiencies (like those with BRCA mutations) cannot effectively repair their DNA, leading to an accumulation of damage and ultimately, cell death – a concept known as "synthetic lethality."
The U.S. Food and Drug Administration (FDA) first approved olaparib, the pioneer PARP inhibitor, in 2014 for advanced ovarian cancer. Since then, other PARP inhibitors like rucaparib and niraparib have gained approval, significantly extending progression-free survival for many patients. However, the initial promise of these drugs has been tempered by the reality that not all patients benefit equally, and a significant proportion eventually develop resistance, underscoring a critical gap in the understanding of their precise mechanisms of action and distribution within complex tumor microenvironments. Globally, ovarian cancer remains the fifth leading cause of cancer death among women, with a 5-year survival rate of approximately 49.7% for all stages, highlighting the urgent need for more effective and durable treatments. Understanding why PARP inhibitors fail in some cases is therefore paramount to improving these outcomes.
Mapping the Invisible: Advanced Imaging Reveals Drug Distribution Discrepancies
The prevailing assumption in drug delivery has often been that if a drug reaches the tumor, it will distribute relatively evenly and exert its effect. This study challenges that notion by providing unprecedented insight into the microscopic world of drug distribution within real human tumor tissue. To achieve this, the researchers utilized thin slices of ovarian tumors, known as "explants," which were maintained alive in the laboratory after being surgically removed from patients. This innovative ex vivo model allowed scientists to treat the samples with PARP inhibitors and directly observe their movement and accumulation in a physiologically relevant context, bypassing the complexities of systemic drug delivery in live organisms while retaining crucial aspects of human tumor biology.
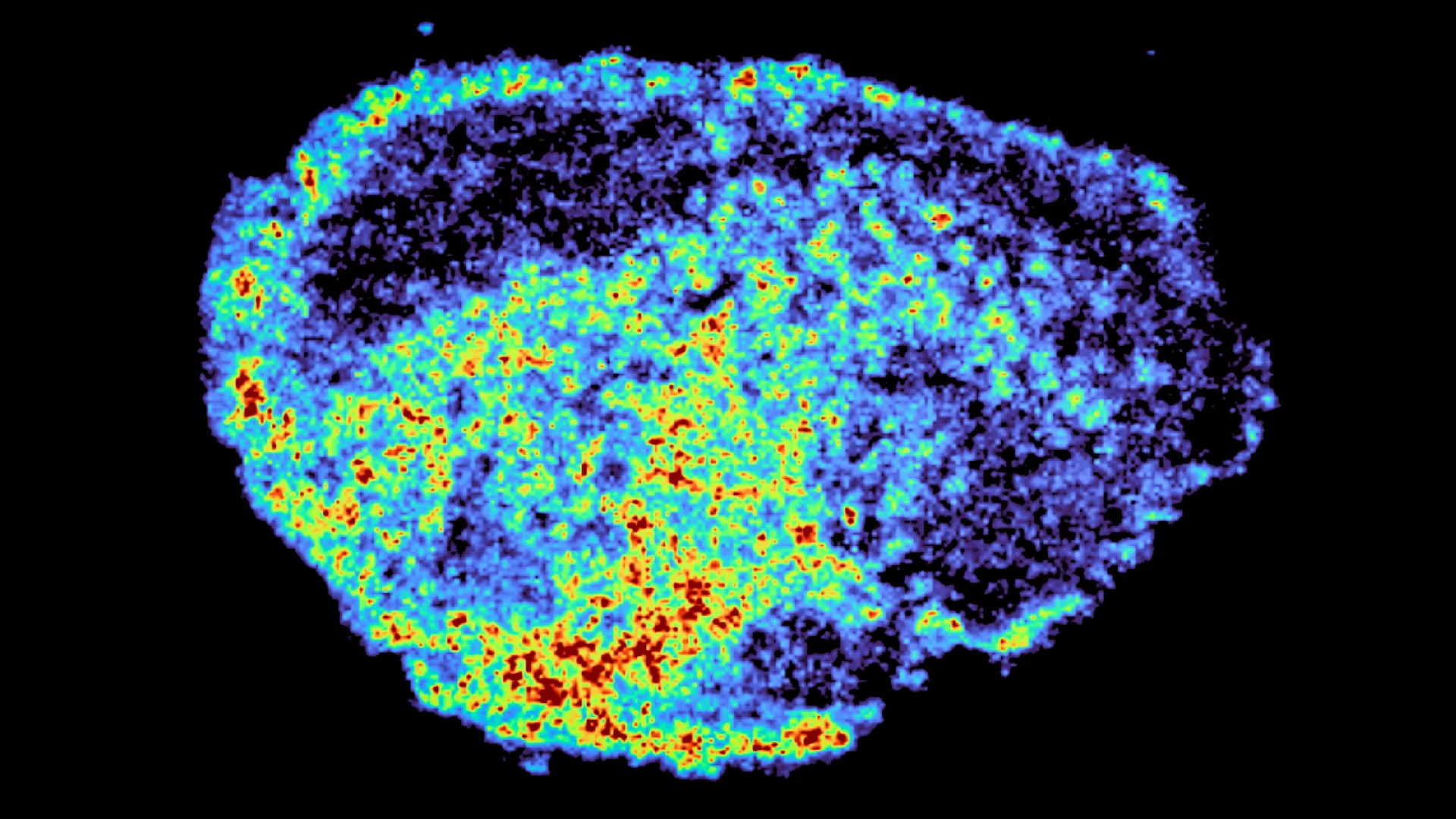
The methodological cornerstone of this research was the synergistic application of two cutting-edge technologies: mass spectrometry imaging (MSI) and spatial transcriptomics. Mass spectrometry imaging provided detailed, high-resolution maps that pinpointed the exact locations where the PARP inhibitor drugs accumulated within the tumor slices. This technique allowed for the direct visualization and quantification of drug molecules at a microscopic level, offering a spatial understanding previously unattainable.
Complementing MSI, spatial transcriptomics was employed to analyze gene activity in specific regions of the same tissue slice that exhibited high or low drug concentrations. By correlating drug presence with gene expression profiles, the researchers could uncover potential biological pathways or cellular characteristics that might influence drug uptake, retention, or metabolism. This powerful combination allowed for a holistic understanding of how drug distribution interacts with cellular biology.
"A novel aspect of this study was the use of mass spectrometry imaging to directly measure and visualize drug uptake in patient tumour tissue. Through the spatial mapping of drug molecules, we could pinpoint regions of high and low drug and compare gene expression, from the same tissue slice, using spatial transcriptomics," explained Dr. Zoe Hall, a senior author of the study and Associate Professor at Imperial’s Department of Metabolism, Digestion and Reproduction. The results were striking, revealing significant variability in drug distribution not only between different patients receiving the same drug dose but also within different regions of a single tumor. This stark heterogeneity underscored the inadequacy of a ‘one-size-fits-all’ approach to drug delivery and monitoring.
Lysosomes: The Hidden Drug Reservoirs Influencing Efficacy
The most profound discovery of the study centered on the role of lysosomes. These small, membrane-bound organelles are ubiquitous within eukaryotic cells and are commonly known as the cell’s "recycling centers." They contain a battery of hydrolytic enzymes that break down waste materials, cellular debris, and foreign substances. However, this research uncovered an unexpected additional function: lysosomes act as significant internal reservoirs for certain PARP inhibitors.
The researchers observed that some PARP inhibitors are actively drawn into these lysosomal compartments, where they become sequestered instead of diffusing evenly throughout the cell’s cytoplasm and nucleus, where their primary targets reside. This sequestration creates internal pockets of high drug concentration within lysosomes, while other parts of the cell, and indeed other cells within the tumor, may experience much lower, potentially sub-therapeutic, drug levels.
Crucially, the study demonstrated that lysosomes do not merely trap the drugs; they act as "slow-release reservoirs." They hold onto the drug and gradually release it over time, effectively increasing the duration of exposure for some cancer cells. While this might sound beneficial, it also means that other cells might be exposed to insufficient drug concentrations, potentially leading to drug resistance. The study also highlighted that not all PARP inhibitors behave identically in this regard. Drugs such as rucaparib and niraparib were significantly affected by this lysosomal sequestration and release mechanism, whereas olaparib, another widely used PARP inhibitor, did not exhibit the same pattern of lysosomal accumulation. This differential behavior among drugs within the same class has critical implications for treatment selection and drug development.
Dr. Carmen Ramirez Moncayo, the first author of the study and a Postdoctoral Researcher at the LMS, articulated the surprise and significance of this finding: "We were surprised to see large variability in drug accumulation at the single-cell level. This variability was driven by the build-up of a drug in lysosomes, which are acting as reservoirs, increasing the exposure of cancer cells to drugs, by storing and releasing the drug when needed." This single-cell variability is a key insight, explaining how even within a seemingly uniform tumor, drug efficacy can differ drastically.
Implications for Personalized Oncology and Future Treatment Strategies
The findings of this study carry profound implications for the future of cancer treatment, particularly for PARP inhibitors, which are already standard of care for several cancers and are under investigation for numerous others, including pancreatic and prostate cancers. A deeper understanding of how these drugs are stored, distributed, and released within cells and tumors could pave the way for more rational, personalized treatment strategies.
For pharmaceutical companies, this research suggests that drug design could be optimized to minimize lysosomal sequestration for drugs where rapid and uniform intracellular distribution is critical, or conversely, to exploit this mechanism for sustained release if deemed beneficial. It also highlights the importance of assessing intracellular drug distribution profiles during preclinical development, beyond just overall tumor uptake.
For clinical oncologists, the study offers a potential avenue for developing predictive biomarkers. If specific molecular signatures or lysosomal characteristics in a patient’s tumor predict uneven drug distribution or significant lysosomal trapping, clinicians might be able to select alternative therapies or adjust dosing regimens to optimize outcomes. This moves closer to the long-held promise of precision medicine, where treatment is tailored to the unique biological characteristics of each patient’s tumor.
"By understanding how drugs are taken up into cells, we can understand whether this influences why cancer drugs work for some people and not for others. Eventually, we hope to be able study the molecular signature of a patient’s tumor to help to tailor therapeutic approaches in a more personalized way," stated Dr. Louise Fets, a senior author of the study and Head of the LMS’ Drug Transport and Tumour metabolism Group. This vision underscores a shift from empirical drug selection to one informed by detailed molecular insights.
Addressing Drug Resistance and Relapse
The uneven distribution of drugs within tumors, exacerbated by lysosomal trapping, also offers a compelling explanation for the development of drug resistance. If certain cancer cells are exposed to sub-lethal concentrations of a drug due to inefficient delivery or sequestration, these cells may survive, adapt, and eventually proliferate, leading to tumor relapse and drug resistance. By understanding the mechanisms that create these drug "cold spots," researchers can devise strategies to ensure more uniform and effective drug delivery, potentially delaying or preventing resistance. This could involve combination therapies, drug reformulations, or interventions that modulate lysosomal function.
Future Trajectories: Bridging Lab to Clinic
While the use of patient tumor explants provided invaluable insights into drug dynamics within human tissue, the study acknowledges its limitations. In real patients, drugs are delivered systemically through the bloodstream, and tumor vasculature is often highly disorganized, forming an additional barrier to uniform drug distribution. This complex physiological environment may further contribute to the heterogeneity observed in drug uptake.
Therefore, future research will build upon these foundational findings by expanding investigations into animal models, which can mimic the systemic drug delivery and tumor microenvironment more accurately. Larger patient cohorts will also be crucial to validate these findings in diverse clinical settings and across different cancer types and stages, including in cases of relapsed cancers where drug resistance is a primary concern. Understanding how tumor structure, blood vessel architecture, and lysosomal storage interact in a living organism will be the next critical step in translating these discoveries into actionable clinical strategies.
This comprehensive research, supported by vital funding from organizations such as the Medical Research Council, Cancer Research UK, a PhD studentship from the Integrative Toxicology Training Partnership administered by the MRC Toxicology Unit, and a Victoria’s Secret Global Fund for Women’s Cancers Career Development Award in partnership with Pelotonia and AACR, represents a significant stride towards unraveling the intricate mechanisms governing cancer drug efficacy. It underscores the importance of continuous fundamental research in pharmacology and cell biology to inform the next generation of precision cancer therapies, ultimately striving for a future where every patient receives the most effective treatment tailored to their unique disease.