Unveiling a "Nuclear Metabolic Fingerprint"
The comprehensive study meticulously documented that different cell types, tissues, and even various cancers exhibit their own unique and distinctive arrangements of these metabolic enzymes within the nuclear confines. These enzyme-DNA interactions form what the researchers have termed a "nuclear metabolic fingerprint," marking the first concrete evidence that human cells possess such unique, metabolism-driven nuclear signatures. This discovery is not merely an observation; it signifies a profound shift in how scientists perceive the compartmentalization and integration of cellular processes.
While the precise roles these nuclear enzymes play are still under investigation, several compelling hypotheses have emerged. They could be directly driving critical chemical reactions essential for nuclear processes, influencing the intricate machinery that switches genes on or off, or even contributing to the structural integrity and organization of the DNA itself. Regardless of their exact functions, these preliminary findings already offer invaluable new insights into the complex mechanisms by which tumors develop, adapt to challenging environments, and, crucially, sometimes resist conventional therapies. The implications extend far beyond basic biology, opening new avenues for understanding and potentially combating cancer.
Dr. Sara Sdelci, a corresponding author of the study and a distinguished researcher at the Centre for Genomic Regulation, emphasized the profound nature of this finding. "Many of these enzymes are fundamental to synthesizing essential building blocks of life, and their nuclear localization is strongly associated with DNA repair mechanisms," Dr. Sdelci stated. "Their unexpected presence in the nucleus may therefore directly shape how cancer cells respond to genotoxic stress, a hallmark characteristic that many chemotherapeutic treatments are designed to induce. It truly represents an entirely new world to explore, brimming with unexplored potential."
The Conventional Divide: Metabolism vs. Genome Regulation
For decades, the fields of metabolism and genome regulation have largely been viewed as distinct and largely separate biological systems. The scientific consensus dictated a clear division of labor: the nucleus, with its meticulously organized chromatin, was understood to be the exclusive domain of the genome, the repository of genetic information, and the command center for gene expression. In stark contrast, metabolic enzymes were primarily understood to operate in the mitochondria, the cell’s powerhouses, where they generate adenosine triphosphate (ATP) through oxidative phosphorylation, and in the cytoplasm, where they manage a vast array of biochemical reactions, from nutrient processing to waste detoxification.
This long-standing assumption, deeply embedded in biological textbooks and research paradigms, underscored the sheer scale of surprise and significance associated with the current discovery. The identification of over 200 metabolic enzymes directly engaged with DNA within the nucleus forces a fundamental re-evaluation of this traditional compartmentalization. It suggests a much more intricate and integrated cellular architecture, where metabolic activity is not just an external energy source but an intrinsic component of nuclear function and genomic integrity.
Methodological Rigor: Isolating Chromatin-Bound Proteins
To achieve this unprecedented identification, the research team employed a sophisticated and robust technique designed to isolate proteins physically attached to chromatin. Chromatin, the natural packaging of DNA within human cells, consists of DNA tightly wound around histone proteins, forming a compact structure that fits the lengthy genetic material into the microscopic nucleus. By specifically targeting and extracting these chromatin-bound proteins, the scientists could precisely identify components that are in direct physical proximity and potential interaction with the genome.
This rigorous methodological approach was applied across an extensive panel of samples, including 44 diverse cancer cell lines and 10 healthy cell types collected from ten distinct human tissues. The sheer breadth of this sampling strategy, encompassing both diseased and healthy states across various physiological contexts, lent significant weight and credibility to the findings. It ensured that the observations were not merely an artifact of a specific cell type or experimental condition but represented a widespread and fundamental biological phenomenon.
The Revelation of a "Mini Metabolism" within the Nucleus
The scale of the discovery was, as the researchers themselves admitted, genuinely surprising. Their meticulous analysis revealed that metabolic enzymes appear to play remarkably active and diverse roles within nuclear biology. Astoundingly, approximately 7 percent of all proteins found physically attached to chromatin turned out to be metabolic enzymes. This significant proportion challenges the notion of the nucleus as an inert genetic vault and instead posits it as a dynamically active compartment with its own unique metabolic landscape.
This observation led the researchers to propose the intriguing concept of a "mini metabolism" operating within the nucleus. This nuclear mini metabolism is not merely a passive presence of enzymes but implies an active network of biochemical reactions taking place directly at the site of the genome. Such a localized metabolic system could provide immediate access to energy, building blocks, or signaling molecules precisely where they are needed for DNA replication, repair, transcription, and other critical nuclear functions, without relying solely on resources shuttled from the cytoplasm or mitochondria. This localized metabolic activity could offer a rapid, efficient, and finely tuned regulatory mechanism for genomic processes.
Surprising Energy Pathways: Oxidative Phosphorylation in the Nucleus
Among the enzymes detected, some were particularly astonishing in their nuclear localization. The team identified proteins involved in oxidative phosphorylation (OXPHOS), the cellular process primarily responsible for generating the vast majority of a cell’s energy in the form of ATP. The presence of OXPHOS enzymes, conventionally confined to the inner mitochondrial membrane, as regular occupants of the nucleus was an unexpected and profound finding. It suggests that the nucleus might possess an independent, albeit perhaps smaller-scale, capacity for generating energy directly on or near the DNA.
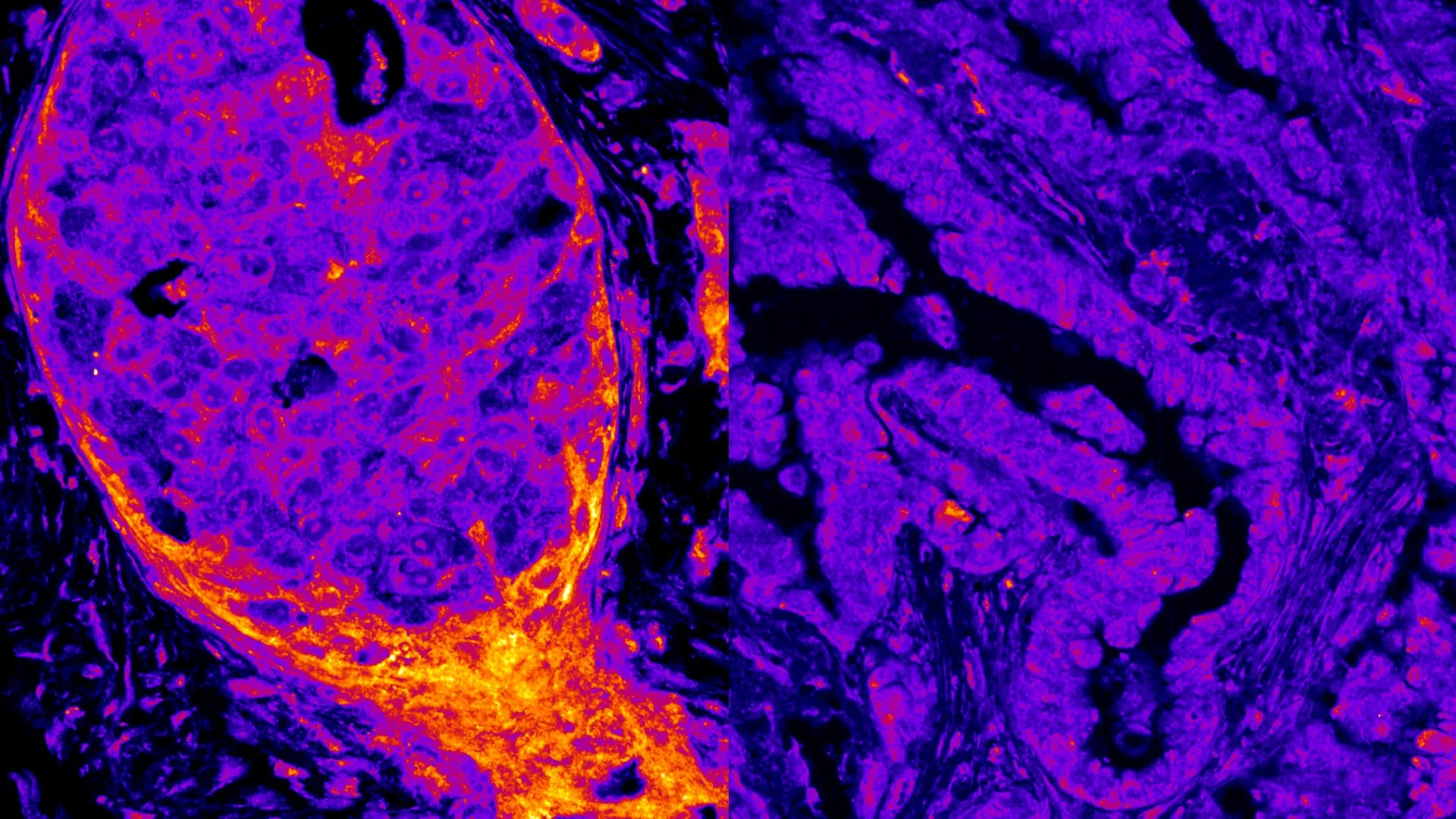
Furthermore, the pattern of these nuclear OXPHOS enzymes was not uniform; it varied significantly depending on the specific cancer type. For instance, oxidative phosphorylation enzymes were commonly observed within the nuclei of breast cancer cells, suggesting a potential role in their unique metabolic adaptations or survival strategies. Conversely, these same enzymes were largely absent or found in much lower concentrations in lung cancer cells. This tissue- and disease-specific variation was not confined to cell lines; when scientists examined actual tumor samples taken directly from patients, they observed the identical trend. This compelling consistency confirmed that nuclear metabolism is highly dynamic and varies depending on the tissue type, the specific disease context, and potentially even the stage or aggressiveness of the cancer.
Dr. Savvas Kourtis, the first author of the study, eloquently summarized the paradigm-shifting nature of these findings: "We’ve traditionally been treating metabolism and genome regulation as two separate universes, operating in isolation from one another. Our work now strongly suggests that they are not only talking to each other but are deeply intertwined. Even more critically, cancer cells might be actively exploiting these intimate conversations and interconnections to survive, proliferate, and resist treatment."
Dynamic Roles: Enzymes Respond to DNA Damage
To move beyond mere observation and begin unraveling the functional significance of these nuclear enzymes, the researchers conducted a series of targeted experiments. They focused their initial efforts on a specific group of enzymes known to be responsible for producing molecules essential for both DNA synthesis and repair. DNA damage is a constant threat to genomic integrity, and robust repair mechanisms are vital for cell survival and preventing mutations that can lead to cancer.
Their experiments yielded compelling results: these particular enzymes exhibited a remarkable ability to gather and concentrate near chromatin specifically when DNA damage occurred. This directed movement and localized accumulation suggest a coordinated response. By concentrating in these vulnerable regions, these enzymes appear to actively assist with the intricate and energy-intensive process of repairing the damaged genome. This finding provides a direct functional link between nuclear metabolism and the maintenance of genomic stability, a critical process for cellular health and disease prevention.
Location-Dependent Functionality: The Case of IMPDH2
One of the most intriguing discoveries revolved around the concept that an enzyme’s function can be profoundly influenced by its subcellular location. The team investigated a specific enzyme known as IMPDH2 (Inosine Monophosphate Dehydrogenase 2). This enzyme displayed strikingly different behaviors and impacts depending on whether it was located in the nucleus or the cytoplasm.
When researchers experimentally manipulated cells to force IMPDH2 to remain exclusively inside the nucleus, it played a crucial role in maintaining genome stability. This suggests a direct nuclear function in protecting and repairing DNA. However, when the very same enzyme was restricted to the cytoplasm, it influenced entirely different cellular pathways, unrelated to genomic integrity. This phenomenon of location-dependent functionality adds another layer of complexity to cellular regulation, indicating that enzymes are not just static catalysts but dynamic players whose roles are contextualized by their environment. Understanding such nuances could be critical for precision medicine and targeted therapies.
Profound Implications for Cancer Research and Therapy
The implications of these findings for cancer research and treatment are substantial and far-reaching. Many current cancer therapies are designed to target specific metabolic processes within cancer cells, aiming to starve them or disrupt their rapid growth. Simultaneously, other therapeutic approaches focus on disrupting DNA repair systems, thereby accumulating damage that becomes lethal to cancer cells. If, as this study strongly suggests, these two fundamental biological processes—metabolism and genome regulation—are far more tightly connected and integrated than previously understood, it necessitates a fundamental re-evaluation of how scientists approach cancer treatment.
This interconnectedness could provide a crucial missing piece in the puzzle of cancer drug resistance. Dr. Sdelci highlighted this potential: "It could help explain why tumors of different origins, even when carrying identical genetic mutations, often respond very differently to chemotherapy, radiotherapy, or targeted inhibitors." The unique nuclear metabolic fingerprints of individual tumors might dictate their susceptibility or resistance to treatments, paving the way for more personalized and effective therapeutic strategies. This paradigm shift could lead to the development of novel combination therapies that simultaneously target both nuclear metabolic pathways and DNA repair, potentially overcoming resistance and improving patient outcomes.
Towards New Diagnostic Biomarkers
Beyond therapeutic implications, this study provides the first large-scale evidence that metabolic enzymes are widely present and functionally relevant inside the nucleus. According to the researchers, systematically mapping where these enzymes are located, understanding their specific functions, and identifying their unique patterns—the "nuclear metabolic fingerprints"—could serve as a powerful new tool for diagnosing cancer. These fingerprints might function as novel biomarkers, allowing for earlier detection, more accurate staging, or even predicting a tumor’s likely response to specific treatments.
Furthermore, identifying these nuclear metabolic pathways could reveal previously unrecognized weaknesses in cancer cells. These vulnerabilities could then be exploited as novel targets for anti-cancer drugs, leading to the development of a new generation of highly specific and effective therapies. The concept of targeting a cell’s "mini metabolism" directly at the genomic level opens up an entirely new therapeutic frontier.
Charting Uncharted Territory: Future Research Directions
Despite the profound nature of these discoveries, the researchers are quick to emphasize that much work remains. This study is merely the opening chapter of a vast new field of inquiry. Scientists still need to definitively determine whether all of the more than 200 enzymes observed in the nucleus are metabolically active, or if some play purely structural or regulatory roles. Furthermore, understanding the specific, unique functions of each individual enzyme within the nuclear context is a monumental task.
Dr. Kourtis underscored this challenge: "Each enzyme may have its own, unique nuclear function, distinct from its cytoplasmic or mitochondrial roles. Therefore, this must be addressed one by one, through meticulous and targeted research." This next phase of research will involve detailed biochemical and genetic studies to elucidate the precise molecular mechanisms at play for each enzyme.
The Mystery of Nuclear Import: How Large Enzymes Cross the Barrier
Another significant and pressing unanswered question involves the fundamental mechanism by which these enzymes gain entry into the nucleus in the first place. The nucleus is meticulously separated from the cytoplasm by a double-layered nuclear envelope, a sophisticated barrier punctuated by nuclear pores. These pores act as selective gates, normally limiting which molecules can pass through, typically based on size and specific transport signals.
Intriguingly, many of the enzymes discovered on DNA are significantly larger than the generally accepted size limits that nuclear pores are believed to accommodate for passive diffusion or even facilitated transport. Despite this apparent size barrier, these bulky proteins still manage to enter the nucleus, suggesting the existence of an as-yet-unknown or highly specialized mechanism for their nuclear import.
Unraveling the intricacies of how this process works—how cells actively transport these large metabolic machines into the nucleus—could eventually reveal precise therapeutic targets. Gaining control over nuclear metabolic activity in diseased cells, particularly cancer cells, by manipulating their nuclear import pathways could represent a powerful and highly targeted strategy for intervention. This ongoing mystery highlights the dynamic and complex nature of cellular organization and the endless frontiers that remain in biological discovery. The findings published in Nature Communications are poised to galvanize a new wave of research, fundamentally altering our understanding of cellular metabolism, genomic regulation, and the intricate dance of life within the human cell.